Case Study (FSA Computational Fellow)
In this guide
In this guide100. As part of its ongoing work to evaluate the potential of BMD modelling in chemical risk assessment, the FSA is working with external and independent collaborators to assess the utility and practicality of the approach in a research setting. The following case study (Carvalho et al 2024 (in prep) (TOX/2023/53) demonstrates how the latest BMD modelling approaches can be applied and integrated to the development and application of a high-throughput gene expression profiling of per- and polyfluoroalkyl substances (PFAS) in primary liver human spheroids. These experiments can help produce toxicologically relevant information for risk assessment by informing read-across in data poor environments.
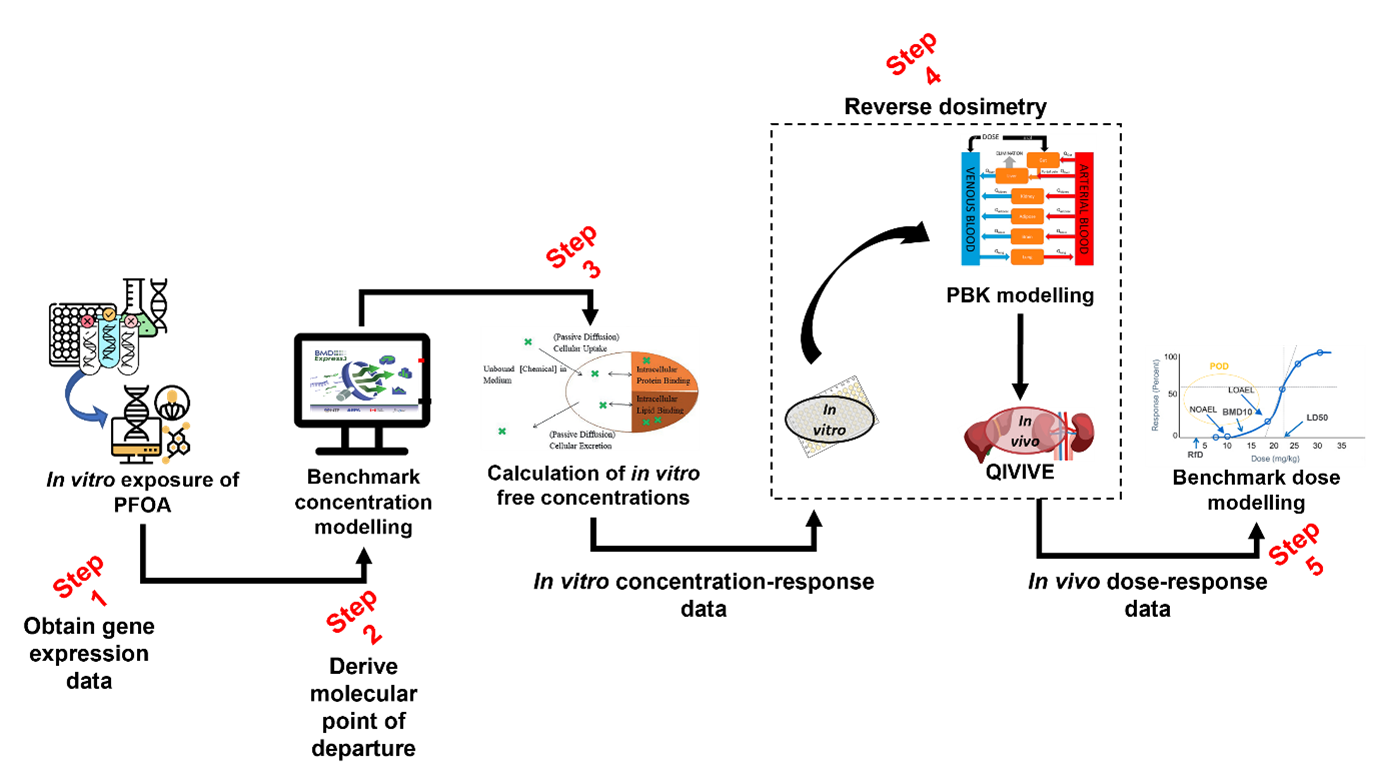
101. Figure 5 provides a summary of the approach and the in-silico workflow used to derive a health-based guidance value for PFAS.
Figure 5. In silico workflow used to derive a health-based guidance value for PFOA.
102. Data was generated in Step 1 (Figure 5) by exposing human liver spheroids to 10 different concentrations of four different PFASs and analysed over four time points: days 1, 4, 10, and 14 (Rowan-Carroll et al. 2021). The expression patterns of about 3000 toxicologically relevant/representative genes were then analysed for their effects in the toxicological assay. The in vitro effects of the PFAS known as perfluorooctanoic acid (PFOA) were modelled at day 10 of exposure, as it was the best time point to observe differentially expressed genes.
103. The BMD software BMDExpress3 Releases · auerbachs/BMDExpress-3 (github.com) (described in the section below) was employed in Step 2 (Figure 5) to derive molecular points of departure from the gene expression data obtained from the literature and Gene Expression Omnibus ( GEO Accession viewer (nih.gov). Modelling was performed under expert guidance. The BMD software provided, as an output, the chosen models for each gene as well as the model averaging for the benchmark concentration as well as both lower and upper bounds of the confidence interval.
104. In silico modelling of the toxicokinetic properties of PFOA is performed as part of the characterisation process using a calibrated model (Step 3 and 4, Figure 5). In the final step (Step 5) of the workflow, PROAST or alternatively, EFSA’s Bayesian BMD modelling suite, (EFSA - Sign in (b2clogin.com) was employed to derive a final health-based guidance value for PFOA using the in vivo dose-response data we had obtained from Step 3 and 4.